Recent genetic studies of complex traits and diseases have focused on the identification of common variants associated with risk through genome-wide association studies (GWAS). Other aspects such as rate of progression, age at onset and the effect of rare variants are generally not investigated. These studies have been very successful in identifying novel loci associated with many complex diseases. The current proposal focuses on these understudied aspects of disease etiology, namely the role of common and rare genetic variation on quantitative diagnostic and prognostic endophenotypes of Alzheimer’s disease (AD). We will use GWAS and exome-chip data to identify single variants, genes and pathways associated with cerebrospinal fluid (CSF) levels of known AD biomarkers (tau, ptau, Aβ, YKL40, VILIP1) and other AD-related proteins (CLU, APOE, TREM2). The integration of these endophenotypes will enable us to disentangle the genetic architecture of AD. With this insight, we will then determine whether those SNPs, genes or pathways are also associated with other AD phenotypes (risk, age at onset or progression), and whether we can use genetic information to increase the diagnostic or prognostic ability of these CSF biomarkers. Further, we will utilize Mendelian Randomization and a novel network-based approach to identify causal plasma and CSF proteins involved in AD and other complex traits. We will have access to a unique resource – a large number of CSF and plasma protein levels – allowing us to leverage unbiased approaches to reveal novel biomarkers and endophenotypes associated with AD and complex traits.
Publications
- Yan Q, Nho K, Del-Aguila JL, Wang X, Risacher SL, Fan KH, Snitz BE, Aizenstein HJ, Mathis CA, Lopez OL, Demirci FY, Feingold E, Klunk WE, Saykin AJ; Alzheimer’s Disease Neuroimaging Initiative (ADNI), Cruchaga C, Kamboh MI. Genome-wide association study of brain amyloid deposition as measured by Pittsburgh Compound-B (PiB)-PET imaging. Mol Psychiatry. 2018; Oct 25. doi: 10.1038/s41380-018-0246-7 PMCID:PMC6219464
- Del-Aguila JL, Fernández MV, Schindler S, Ibanez L, Deming Y, Ma S, Saef B, Black K, Budde J, Norton J, Chasse R; Alzheimer’s Disease Neuroimaging Initiative (ADNI), Harari O, Goate A, Xiong C, Morris JC, Cruchaga C. Assessment of the Genetic Architecture of Alzheimer’s Disease Risk in Rate of Memory Decline. J Alzheimers Dis. 2018; 62(2):745-756. PMCID: PMC5989565
- Huang KL, Marcora E, Pimenova AA, DiNarzo AF, Kapoor M, Jin SC, Harari O, Bertelsen S, Fairfax BP, Czajkowski J, Kauwe JSK, Cruchaga C, Hao K, Goate AM. A common haplotype lowers PU.1 expression in myeloid cells and delays onset of Alzheimer’s disease. Nat Neuroscience 2017. 20(8):1052-1061. PMCID: PMC5759334
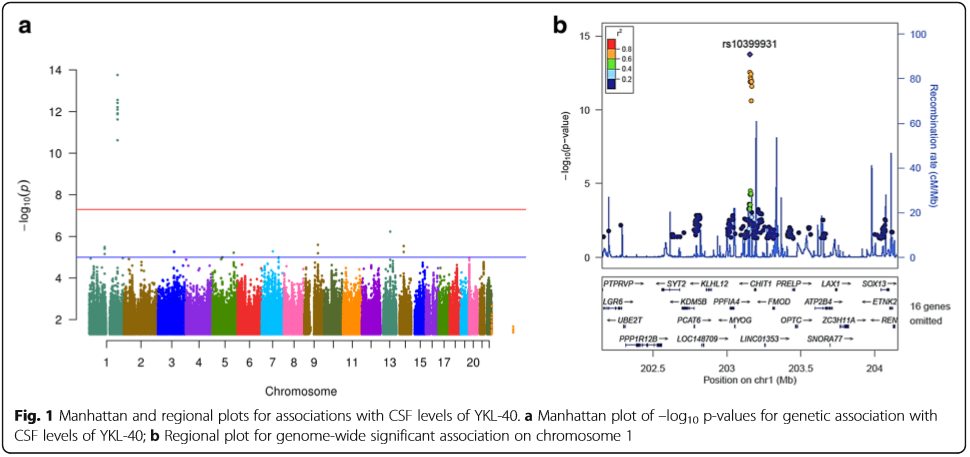
- Deming Y, Li Z, Kapoor M, Harari O, Del-Aguila JL, Black K, Carrell D, Cai Y, Fernandez MV, Budde J, Ma S, Saef B, Howells B, Huang KL, Bertelsen S, Fagan AM, Holtzman DM, Morris JC, Kauwe JS, Goate AM, Cruchaga C. Genome-wide association study identifies four novel loci associated with Alzheimer’s endophenotypes and disease modifiers. Acta Neuropathol. 2017; 133(5):839-856. PMCID: PMC5613285
- Piccio L, Deming Y, Del-Águila JL, Ghezzi L, Holtzman DM, Fagan AM, Fenoglio C, Galimberti D, Borroni B, Cruchaga C. Cerebrospinal fluid soluble TREM2 is higher in Alzheimer disease and associated with mutation status. Acta Neuropathol 2016; 131(6):925-33. PMCID: PMC4867123
- Deming Y, Xia J, Cai Y, Lord J, Holmans P, Bertelsen S, Holtzman D, Morris JC, Bales K, Pickering EH, Kauwe J, Goate A, Cruchaga C; Alzheimer’s Disease Neuroimaging Initiative (ADNI). A potential endophenotype for Alzheimer’s disease: cerebrospinal fluid clusterin. Neurobiol Aging 2016; 37:208. PMCID: PMC5118651
- Kauwe JS, Bailey MH, Ridge PG, Perry R, Wadsworth ME, Hoyt KL, Staley LA, Karch CM, Harari O, Cruchaga C, Ainscough BJ, Bales K, Pickering EH, Bertelsen S; the Alzheimer’s Disease Neuroimaging Initiative, Fagan AM, Holtzman DM, Morris JC, Goate AM. Genome-Wide Association Study of CSF Levels of 59 Alzheimer’s Disease Candidate Proteins: Significant Associations with Proteins Involved in Amyloid Processing and Inflammation. PloS Genet 2014; 10(10):e1004758. PMCID: PMC4207667.
- Cruchaga C, Ebbert MT, Kauwe JS. Genetic discoveries in AD using CSF amyloid and tau. Curr Genet Med Res 2014; 2(1):23-29. PMCID: PMC3979575.
- Benitez BA, Jin SC, Guerreiro R, Graham R, Lord J, Harold D, Sims R, Lambert JC, Gibbs JR, Bras J, Sassi C, Harari O, Bertelsen S, Lupton MK, Powell J, Bellenguez C, Brown K, Medway C, Haddick PC, van der Brug MP, Bhangale T, Ortmann W, Behrens T, Mayeux R, Pericak-Vance MA, Farrer LA, Schellenberg GD, Haines JL, Turton J, Braae A, Barber I, Fagan AM, Holtzman DM, Morris JC; The 3C Study Group, the EADI consortium, the Alzheimer’s Disease Genetic Consortium (ADGC), Alzheimer’s Disease Neuroimaging Initiative (ADNI), the GERAD Consortium, Williams J, Kauwe JS, Amouyel P, Morgan K, Singleton A, Hardy J, Goate AM, Cruchaga C. Missense variant in TREML2 protects against Alzheimer’s disease. Neurobiol Aging 2014; 35(6):1510. PMCID: PMC3961557
- Cruchaga C, Kauwe JSK, Harari O, Jin SC, Cai Y, Karch CM, Benitez B, Jeng AT, Skorupa T, Carrell D, Bertelsen S, Bailey M, McKean D, Shulman JM, De Jager PL, Chibnik L, Bennett DA, Fagan AN, Holtzman DM, Morris JC, the Alzheimer Disease Genetic Consortium (ADGC), Alzheimer’s Disease Neuroimaging Initiative (ADNI), the GERAD Consortium, and Alison M. Goate. GWAS of cerebrospinal fluid tau levels identifies novel risk variants for Alzheimer’s disease. Neuron 2013; 78(2):256-68. PMCID: PMC3664945